


A large-scale and diverse dataset of raw musculoskeletal MRI
MosaicMRI is the largest and most diverse open-source raw musculoskeletal MRI dataset to date.
It spans 10 anatomies, 2,671 volumes, and 80,156 slices.
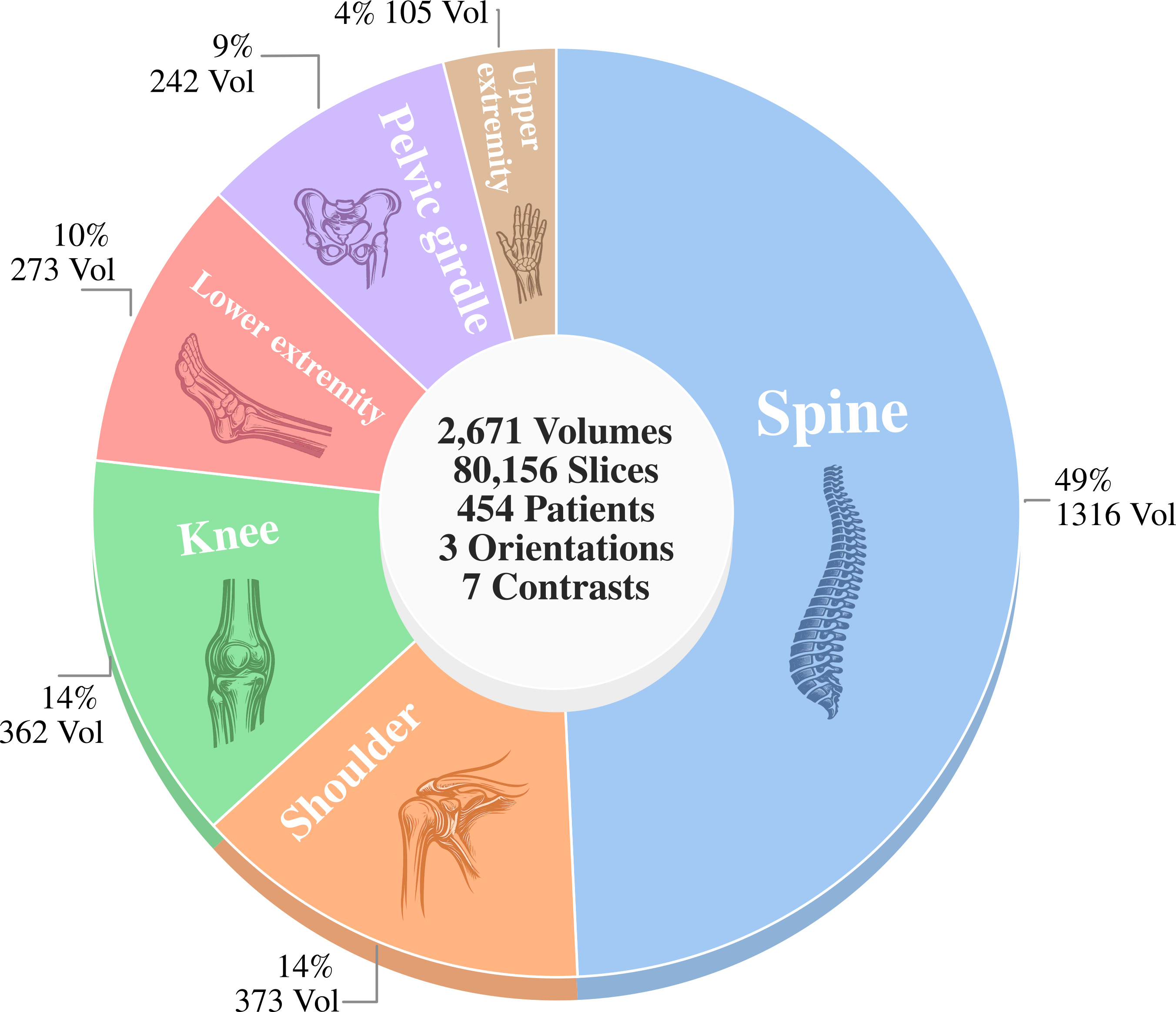
MosaicMRI brings realistic clinical variability across anatomy, contrast, orientation, and coil configuration far beyond the existing brain- and knee-focused datasets.
MosaicMRI fascilitates research on accelerated MRI, low-field reconstruction, motion suppression, and other real-world reconstruction challenges.
This initial release goes beyond typical accelerated reconstruction to probe anatomical and contrast generalization under real-world variability.
MosaicMRI provides a testbed for studying key foundation model challenges, including scaling laws, data synergies, continual learning, data mixtures, reliability, and out-of-distribution generalization.
Orientation-specific examples grouped by anatomy. Select a category to view axial, coronal, and sagittal samples.
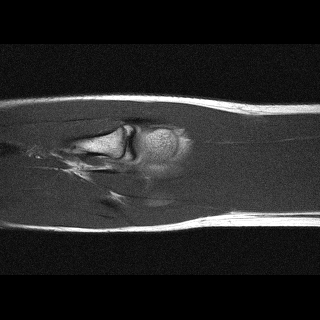
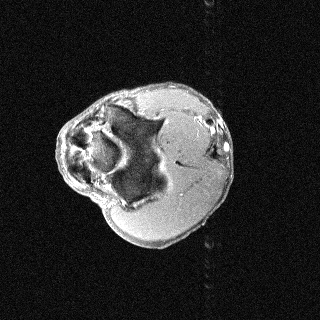
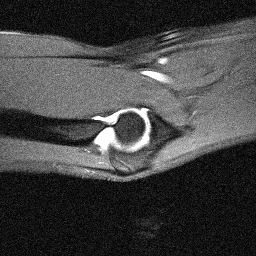
MosaicMRI is designed for learning-based MRI under realistic clinical variability in anatomy, contrast, orientation, and coil configuration.
Data were collected on a 1.5T Siemens Magnetom Avantofit scanner between July 15, 2025 and September 23, 2025. We removed incomplete exams, localizers/planning scans, calibration-only acquisitions, and protocols not suited for slice-based reconstruction.
Remaining scans were visually quality-checked and stored as HDF5 with ISMRMRD-compatible headers and fastMRI-style internal layout.
AX/SAG/COR), coarse contrast, fat-suppression flag, and anatomical category.Splits are patient-disjoint to avoid leakage, with target ratios 70% train, 15% val, and 15% test. Assignment was optimized to balance slice counts while preserving per-anatomy coverage across splits.
| Split | Scans | Patients | Slices |
|---|---|---|---|
| train | 1,873 | 303 | 56,235 |
| val | 398 | 68 | 12,027 |
| test | 400 | 79 | 11,894 |
File organization and baseline usage for reconstruction experiments.
Directory layout (current release statistics):
MosaicMRI/
multicoil_train/ (1,744 files, 2,381.92 GiB)
*.h5
multicoil_val/ (398 files, 579.77 GiB)
*.h5
multicoil_test/ (64 files, 71.58 GiB)
*.h5
anatomy_transfer_challenge/
ankle/ (20 files, 49.40 GiB)
*.h5
contrast_generalization_challenge/
T1_FS/ (17 files, 20.74 GiB)
*.h5multicoil_train, multicoil_val, and multicoil_test are the standard reconstruction splits; multicoil_test contains both 4x and 8x accelerated test inputs.anatomy_transfer_challenge/ankle and contrast_generalization_challenge/T1_FS are challenge-specific evaluation subsets.ismrmrd_header, kspace, and reconstruction_rss.tStudyDescription and tProtocolName).A helper script in the GitHub repository reads these metadata fields and plots one reference slice.
Minimal steps to download a file, apply a mask, and run a baseline reconstruction.
git clone https://github.com/AIF4S/mosaicmri
cd mosaicmri
conda env create -f varnet/environment.yml
conda activate mosaic_mri_varnetpython varnet/run_pretrained_varnet_inference.py --state_dict_file /path/to/weights.ckpt --data_path /path/to/MosaicMRI/multicoil_test --output_path /path/to/recons --accelerations 8 --center_fractions 0.04To obtain access, submit the request form below. Approved users receive access through the gated Hugging Face dataset.
A three-track, 8x-acceleration benchmark: Mixed Anatomy Reconstruction, Anatomy Generalization (held-out ankle), and Contrast Generalization (held-out T1-FS). Participants upload reconstructed H5 files and are evaluated against hidden ground truth with PSNR, SSIM, and NMSE leaderboards.
Go to BenchmarkAccess is granted for research use after manual review.
MosaicMRI is released for non-commercial research and method development under the posted license terms.
Metadata is de-identified before release.
Please cite the dataset paper if you use MosaicMRI.
@article{mosaicmri_2026,
title = {MosaicMRI: A Diverse Dataset and Benchmark for Raw Musculoskeletal MRI},
author = {Arguello, Paula and Tinaz, Berk and Mohammad, Shahab Sepehri and Soltanolkotabi, Maryam and Soltanolkotabi, Mahdi},
journal = {arXiv},
year = {2026},
doi = {10.48550/arXiv.2604.11762}
}Citation metadata will be updated if publication details change.



University of Utah

University of Southern California (USC)

University of California, Irvine (UCI)